Bengaluru: Researchers have reported the design of a new class of synthetic peptides that can not only block the entry of SARS-CoV-2 virus entry into cells, but also clump the virions (virus particles) together, reducing their ability to infect. This novel approach provides an alternative mechanism to render viruses like SARS-CoV-2 inactive, promising a new class of peptides as antivirals.
The rapid emergence of new strains of the SARS-CoV-2 virus has diminished the protection offered by COVID-19 vaccines calling for new approaches to prevent infection by the virus.
It is known that protein-protein interaction is often like that of a lock and a key. This interaction can be hampered by synthetic peptide that mimics, competes with, and prevents the ‘key’ from binding to the ‘lock’, or vice versa.
Scientists at the Indian Institute of Science (IISc), in collaboration with researchers from the CSIR-Institute of Microbial Technology, have exploited this approach to design peptides that can bind to and block the spike protein on the surface of SARS-CoV-2 virus. This binding was further characterised extensively by cryo-electron microscopy (cryo-EM) and other biophysical methods.
The research was supported under the COVID-19 IRPHA call by Science and Engineering Research Board (SERB), a statutory body of Department of Science and Technology (DST).
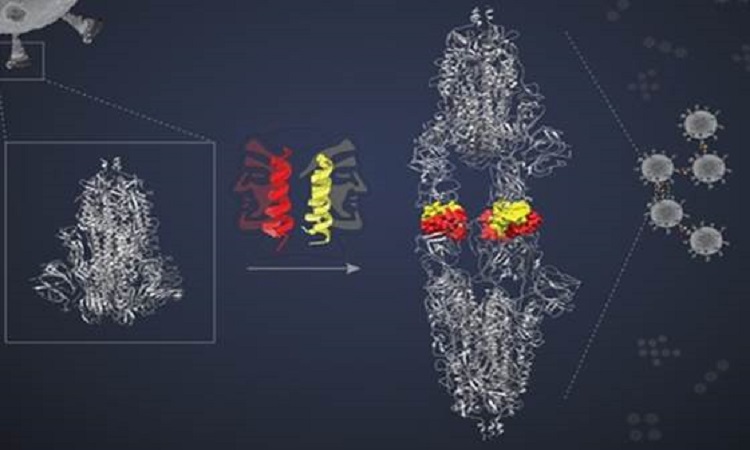
The designed peptides are helical, hairpin-shaped, each capable of pairing up with another of its kind, forming what is known as a dimer. Each dimeric ‘bundle’ presents two ‘faces’ to interact with two target molecules. In the study published in Nature Chemical Biology, the researchers hypothesised that the two faces would bind to two separate target proteins locking all four in a complex and blocking the targets’ action.
Also Read: People risk transmitting Covid19 while talking, finds study
The team decided to test their hypothesis by using a peptide called SIH-5 to target the interaction between the Spike (S) protein of SARS-CoV-2 and ACE2 protein, the SARS-CoV-2 receptor in human cells.
The S protein is a trimer – a complex of three identical polypeptides. Each polypeptide contains a Receptor Binding Domain (RBD) that binds to the ACE2 receptor on the host cell surface. This interaction facilitates viral entry into the cell.
The SIH-5 peptide was designed to block the binding of the RBD to human ACE2. When a SIH-5 dimer encountered an S protein, one of its faces bound tightly to one of the three RBDs on the S protein trimer, and the other face bound to an RBD from a different S protein. This ‘cross-linking’ allowed the SIH-5 to block both S proteins at the same time. Under cryo-EM, the S proteins targeted by SIH-5 appeared to be attached head-to-head, and the spike proteins were being forced to form dimers. Subsequently, the researchers showed that SIH-5 inactivated the viruses efficiently by cross-linking the spike proteins from different virus particles.
Also Read: Amid Covid resurge in Jharkhand, RIMS starts Genome sequencing
The team consisting of B Khatri, I Pramanick, SK Malladi, RS Rajmani, P Ghosh, N Sengupta, R Varadarajan, S Dutta and J Chatterjee from Indian Institute of Science (IISc), R Rahisuddin, S Kumar, N Kumar, S Kumaran and RP Ringe from CSIR-Institute of Microbial Technology, tested the peptide for toxicity in mammalian cells in the lab and found it to be safe. When hamsters were dosed with the peptide and subsequently exposed to a high dose of SARS-CoV-2, they showed decreased viral load as well as much less cell damage in the lungs compared to hamsters exposed only to the virus, demonstrating the promise of this class of peptides as antivirals.
The researchers believe that with minor modifications and peptide engineering, this lab-made miniprotein could inhibit other protein-protein interactions as well.